Project Description
This project continues with specific application to the National COVID Cohort Collaborative (N3C) initiative.
A sandbox is an isolated testing environment that enables users to run programs or execute files without affecting the application, system, or platform on which they run. The sandbox allows developers to test programming code for optimal use of the tool.
The ability to share and integrate clinical phenotype data in support of clinical and translational research requires a systematic approach to evaluating and reporting upon data quality. In response to such needs, this sandbox project will establish a cloud-based sandbox environment in which CTSA hubs can develop, evaluate, and share tools and methods for data quality assessment. It will include a pilot that leverages the Accrual to Clinical Trials (ACT) Network data to understand the quantity and completeness of ACT data and differences in coding practices across institutions. Our objectives in doing so are to: (1) reduce redundancies in such efforts and increase economies-of-scale across the CTSA network, (2) ensure the reproducibility and rigor of such data quality assessment tools and methods, and (3) expedite access to “best-of-breed” data quality assessment tool and methods by all CTSA network participants and partners. The project has three specific aims:
- To create a cloud-based environment that can enable the systematic verification and validation of semi-automated and/or automated clinical data assessment tools and methods
- To populate the “data quality sandbox” with necessary and appropriate reference data sets to be used in shared verification/validation tasks, and further, design and implement a data permutation engine that can be used to systematically introduce data quality relevant errors or features to the reference data sets to enable appropriate performance testing paradigms
- To demonstrate the “data quality sandbox” by engaging a group of CTSA hubs to contribute semi-automated and/or automated clinical data assessment tools and methods to the project and demonstrate their performance, reproducibility, and rigor in such a shared environment.
Simultaneously, the project will continuously generate best practice and technical documentation to support and enable the adoption and adaptation of all of the above work products and technical artifacts.
Critical to the success of our project will be the engagement of a broad and representative group of CTSA hubs who can participate both in the design of the “data quality sandbox” as well as adopt or adapt the tools and methods that will be verified, validated, and shared via this mechanism. We will seek to achieve such engagement and participation via multiple approaches, including: 1) direct interaction with the CTSA network informatics community-of-practice; 2) the establishment of a series of interactive webinars to introduce the sandbox environment and its capabilities to the CTSA network; and 3) the adoption of FAIR principles to guide all aspects of the project so as to ensure the rapid shareability of all research products and knowledge generated therein.
Figure 2 below shows a high-level information architecture model for the planned “data quality sandbox” environment:
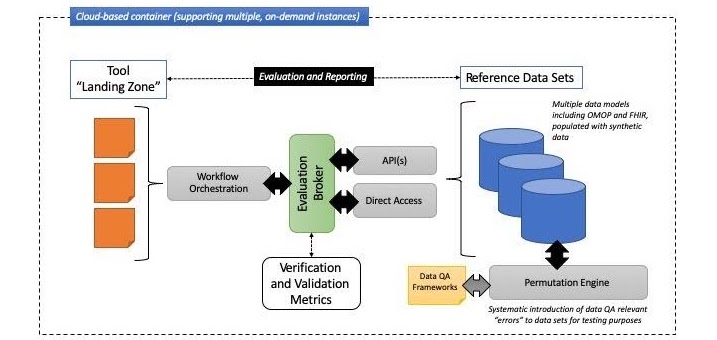
Figure 2: information architecture model for the planned "data quality sandbox" environment
This project will be evaluated using a combination of process (users, access to the sandbox, number of tools/methods and reference data sets being contributed to the environment) and outcomes measures (new tools that are verified/validated and subsequently adopted by other CTSA hubs, new studies or research programs that have generated demonstrable outcomes and that utilized tools/methods developed and demonstrated in the sandbox environment).


